CHAMP (Convex Hull of Admissible Modularity Partitions)¶
A modularity based tool for screening a set of partitions.
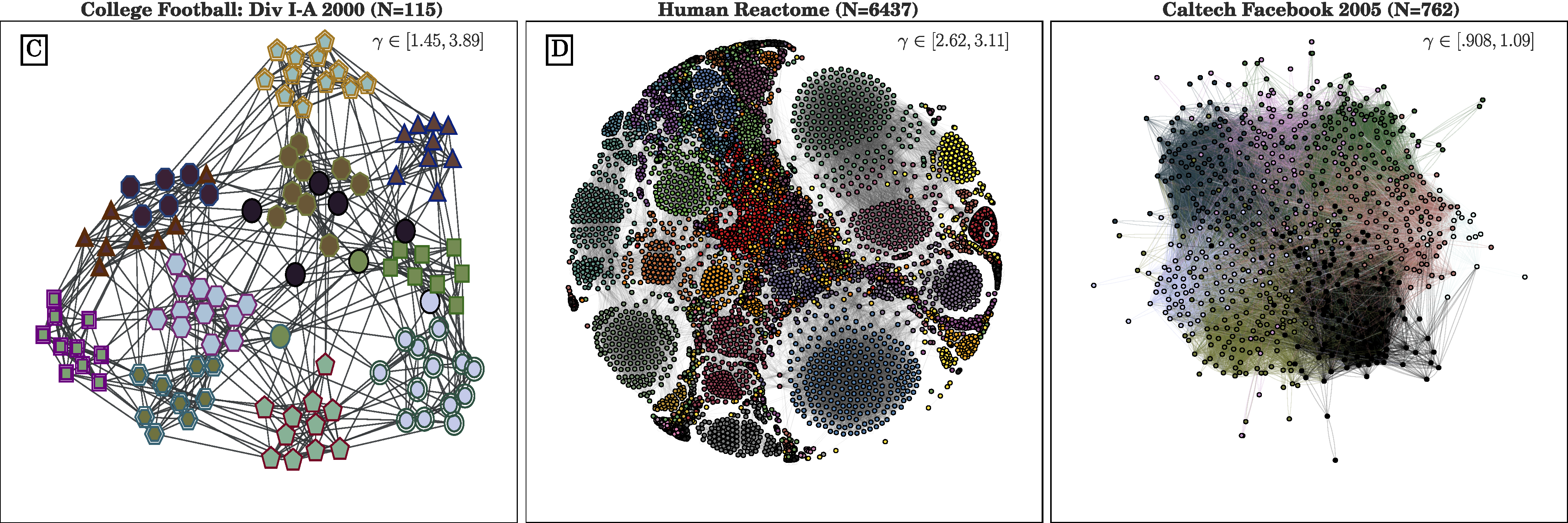
The CHAMP python package provides two levels of functionality:
- Identifying the subset of partitions from a group of partitions (regardless of how they were discovered) with optimal modularity. See Running CHAMP.
- Parallelized implementation of modularity based community detection method, louvain with efficient filtering (ala CHAMP), management, and storage of the generated partitions. See Louvain Parallel Extension .
Contents:¶
Download and Installation:¶
The CHAMP module is hosted on PyPi. The easiest way to install is via the pip command:
pip install champ
For installation from source, the latest version of champ can be downloaded from GitHub:
For basic installation:
python setup.py install
We have also created a conda-forge recipe for creating a conda environment. If you wish to install through conda, first add conda-forge channel:
conda config --add channels conda-forge
Then you can create a environment using prepackaged versions from conda forge:
conda install champ
Dependencies¶
Most of the dependencies for CHAMP are fairly standard tools for data analysis in Python, with the exception of louvain_igraph. They include :
- NumPy : Python numerical analysis library.
- sklearn :Machine learning tools for python.
- python-igraph :igraph python version for manipulation of networks.
- matplotlib :Python data visualization library.
- louvain :Vincent Traag’s implementation of louvain algorithm.
- h5py : HDF5 file format library for python.
These should all be handled automatically if using pip to install.
Citation¶
Please cite:
| [1] | William H. Weir, Scott Emmons, Ryan Gibson, Dane Taylor, and Peter J. Mucha. Post-processing partitions to identify domains of modularity optimization. Algorithms, 2017. URL: http://www.mdpi.com/1999-4893/10/3/93, doi:10.3390/a10030093. |
| [2] | William H. Weir, Ryan Gibson, and Peter J Mucha. Champ package: convex hull of admissible modularity partitions in python and matlab. 2017. https://github.com/wweir827/CHAMP. |
For more details and results see our manuscript
Acknowledgements¶
This project was supported by the James S. McDonnell Foundation 21st Century ScienceInitiative-Complex Systems Scholar Award grant #220020315, by the National Institutes of Health through Award Numbers R01HD075712, R56DK111930 and T32GM067553, and by the CDC Prevention Epicenter Program. The content is solely the responsibility of the authors and does not necessarily represent the official views of the funding agencies.